Single-molecule sequencing (Single-molecule Sequencing, SMS) is a third-generation sequencing technology developed based on the first-generation Sanger sequencing and the second-generation NGS high-throughput sequencing technology. Because the DNA molecules are sequenced without PCR amplification, and the separate sequencing of each DNA molecule is named. Helico Bioscices Company launched the first single molecule sequencing platform in the world, HeliScope, in 2008 based on synthetic sequencing theory, but the average read length of the sequencing is relatively short, and the overall sequencing error rate of the system is high [1]. After that, single-molecule long-read long sequencing technology appeared. At present, the commercial long-read long sequencing technology mainly includes single-molecule real-time sequencing technology (Single-molecule Real-time, SMRT) of Pacific Biosciences (PacBio) and Oxford Nanopore (Oxford Nanopore Technologies, ONT) of [2]. SMS and SMRT techniques mainly convert four different bases into fluorescent signals, and then distinguish the bases by transforming the amplified signal. The nanopore sequencing technology is to convert four bases into electrical signals, and then collect, sort out, transform and make data output, which is quite different from the signal processing of single-molecule fluorescence sequencing technology. Therefore, nanopore sequencing technology is also known as the fourth generation sequencing technology. This paper will mainly introduce the principles and characteristics of nanopore single molecule sequencing technology and its application in the field of in vitro diagnosis.
1、Principles of Nanopore sequencing
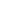
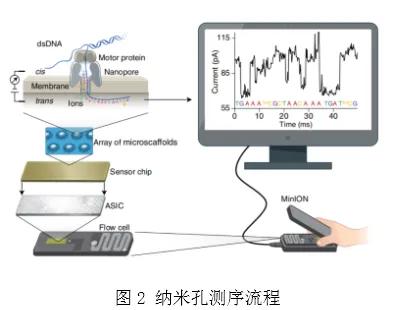
Nanopore single-molecule sequencing technology is based on electrical signal sequencing technology. Compared with other sequencing technologies, the sample processing of nanopore sequencing technology is extremely simple, without DNA polymerase or ligase, nor dNTPs. The sequencing process contains the following key substances, as shown in Figure 1 and Figure 2.

Nanopins (Nanopore), transmembrane proteins that can be embedded in a cell membrane as ions or molecular channels, with natural protein nanopore.
Polymer film (Membrane), the transmembrane protein is embedded in a highly resistivity synthetic polymeric membrane, the membrane is flanked by an ionic solution, and when different potentials are added on both sides, the ions will flow in the pore, forming a current.
Motor protein (Motor protein), for nanopore sequencing libraries, is required to be attached to the connector to push DNA or RNA molecules into the nanopore.
Connect arm (Tether) to anchor DNA or RNA strands, prevent it from drifting in solution and bring it into the nanopore.
The synthetic multimeric membrane is immersed in the ionic solution, and the membrane is covered with the transmembrane channel protein nanopore composed of helicase and protein pore, and different voltages are applied on both sides of the membrane to form voltage differences. Because the polymer membrane is not conductive, the current conducts only through the nanopores. The connecting arm guides the chain to be tested into the nanopore, and then uncoiled under the traction of the motor protein (Figure 1). Different bases pass through the constant electric field of the nanopore to cause different changes of current. Computer software and artificial intelligence algorithms are used to identify and infer the base type, thus completing the sequencing [3] of DNA or RNA (see Figure 2).
2、Type of the nanopore
1. The biological nanopore sp
The concept of nanopore sequencing was first proposed in the 1980s, and has been realized with the technological development of nanopore proteins and motor proteins. The first nanopore protein found to influence ionic currents through DNA or RNA and to enable it to be detected was the α -hemolysin protein [4] proposed in 1996 by Kasianowicz et al. However, the rate of DNA through the nanopoore is too fast to obtain an effective current signal. Later, in 2010, M. smegmatis protein A (Msp-A) nanopore were found to slow the passage of DNA through the nanopore and improve the detection sensitivity of DNA single base, [5]. Subsequently, in 2014, researchers found that using the phi29 DNA polymerase as the motor protein [6], it could control the speed of DNA passing through the nanopore.
Biological nanopores mainly contain the above three kinds, all of which are natural nanopores formed by a certain protein molecule embedded on the phospholipid membrane, which can undergo flexible biochemical modification. However, the problems of membrane stability, current noise and other aspects of biological nanopore limit its development to some extent. Oxford Nanopore company has made some progress in the application of protein nanopore, and their GridION and MinION systems are sequencing platforms based on biological protein nanopore.
2. Solid-state nanopore s
In 2001 and started the solid-state nanopore study [7], Li et al. Solid state nanopore mainly in silicon oxide, graphene and other solid state material by ion beam etching technology of nanopore, because of its size adjustable, high reliability, easy to modify, is widely used in DNA sequencing, protein detection and energy conversion research [8], which is more common used for DNA detection of solid state nanopore is silicon nitride nanopore and graphene nanopore.
Compared with biological nanopres, solid-state nanopres have significant advantages in stability, current noise and process integration. However, due to the limited level of semiconductor process manufacturing, the manufacturing of solid-state nanopres is relatively complex and expensive.
3、Characteristics of Nanopore sequencing
Compared with other sequencing platforms, nanopore sequencing, as a new sequencing technology, has different characteristics in terms of cost, speed, read length and accuracy. Its advantages are very significant, which can be summarized as follows:
1.for the longer sequencing read length
Nanopore sequencing technology uses the change of electrical signal when the base passes through the nanopore. Theoretically, all the nucleic acid sequence passing through the nanopore can be detected, and the read length is only limited by the length of the measured single-stranded DNA [9]. Lenglength can provide a more complete and continuous genome assembly, with significant advantages in genomes with large structural variation and high-level repeat regions.
2. Fast and real-time sequencing is available
Compared with other traditional sequencing, nanopore sequencing achieves dynamic real-time sequencing in a real sense, and can output results. Users can understand the quality and status of samples in the early stage of sequencing, or stop sequencing after obtaining enough data. At the same time, the time required by nanopore sequencing technology is much shorter than that of other sequencing methods, which saves the operation time and reduces the sequencing cost.
3.The Nanopore sequencing device is simple and portable
At present, the most mature MinION sequencer is only the length of a pen, weighing about 100g and only 10cm. It can be inserted with high-speed USB into the computer, with its very small volume completely overturned peoples impression of the sequencer, known as "U disk sequencer" [10]. Due to its portability, real-time sequencing can be completed in various complex environments such as laboratory, field and even space, which can ensure the timeliness of emergency epidemic treatment.
4. The RNA can be sequenced directly
Nanopore sequencing can directly sequence all kinds of raw DNA and RNA, which not only saves time and cost, and retains the information of base modification completely, but also avoids the bias and possible introduced mutations generated by the reverse transcription of RNA into DNA amplification. The workflow of its library preparation is also relatively simple.
Although the advantages of nanopore sequencing are very obvious, compared with previous generations of technology in terms of cost and speed, but it is still in the initial stage, and there are many problems from the sequencing principle to the manufacturing process. First, the detection accuracy is relatively low, which is a problem that has been solved in the development of nanopore sequencing. In addition to optimizing the nanopore and motor proteins, it is also solved by independently researched and developed algorithms. Second, the detection flux and the library yield also limit the application of this technique. Standardized procedures for high-yield library preparation and sequencing for small numbers of samples are still lacking. And it is not always possible to obtain enough large and complete high-molecular-weight DNA and full-length RNA from clinical samples. A trade-off between read length and yield.
4、Application of Nanopore sequencing
Nanopore sequencing has unique advantages: long length and fast sequencing speed, which gives it a great advantage in some scenarios, but the low throughput, high price and low accuracy also severely limit its application scenarios. According to previous analysis, application scenarios that require high read length, but low sequence number and accuracy are the optimal scenarios for nanopore sequencing.
1. Large genome splicing
In the previous genome splicing based on short fragments, some animal and plant genomes are extremely polyploid, highly repetitive and highly heterozygous properties. Nanopore sequencing technology has the characteristics of long read and long, which is conducive to the splicing of large genomes and can greatly improve the integrity of the genome.
2. and the full-length transcriptome
Previous transcriptome analysis failed to direct RNA sequencing, which often required to interrupt mRNA first and then reverse transcribe into cDNA, unable to obtain and analyze full-length transcripts. The long read length of nanopore can accurately identify multiple homologous ers of each gene, simple and accurate; and can sequence RNA directly to identify the base modification of RNA.
3. Structural variation of large segments
The genome will produce many large segments of structural variations related to human diseases (such as deletion, inversion and translocation), short sequencing read length cannot accurately detect these variants, and the read length of nanopore sequencing is longer, suitable for the detection of large segment structural variation, which has a good development prospect in disease research and other aspects.
4. Rapid identification of [11] by pathogenic microorganisms
Due to the real-time and fast nanopore sequencing, the sequencing can be directly sequenced at the acquisition point, and the sequence information is obtained in real time for species classification and identification to complete the rapid identification of pathogenic microorganisms. At present, the application of nanopore sequencing technology in the rapid detection of infectious diseases and clinical infections is relatively extensive.
5、Conclusions and outlook
Nanopore sequencing technology has developed rapidly in recent years. Compared with other sequencing technologies, it has been widely used in various fields with its characteristics such as ultra-long and long reading, real-time monitoring, simplicity and portability, but the nanopore sequencing technology is not perfect at present. At present, many domestic and foreign enterprises are in the process of preparing related single molecule sequencing products, mainly focusing on the direction of pathogen detection and early screening of tumor and genetic diseases. In the future, we will continue to pay attention to the accuracy of related products, the detection flux and the related technical progress of library preparation.
reference documentation::
[1] Steinmann KE,Hart CE,Thompson JF,Milos PM. Helicos Single-molecule Sequencing of Bacterial Genomes[J]. High-Throughput Next Generation Sequencing,2011,733:3–24.
[2] Maitra RD,Kim J,Dunbar WB. Recent Advances in Nano-pore Sequencing[J]. Electrophoresis,2012,33 (23):3418–3428.
[3] Wang Y, Zhao Y, Bollas A,etal. Nanopore sequencing technology, bioinformatics and applications[J]. Nature Biotechnology,2021,39(11):1348-1365.
[4] Kasianowicz JJ. Some Physics and Applications for DNA TransportThrough Single Nanopores. APS Meeting Abstracts, 2002.
[5] Butler TZ,Pavlenok M,Derrington IM, etal.Single-molecule DNA detection with an engineered MspA protein manopore[J].Proc Natl Acad Sci USA,2008,105(52):20647-20652.
[6] Manrao EA,Derrington IM,Laszlo AH,etal.Reading DAN at single-nucleotide resolution with a mutant MspA manoproe and phi29 DNA polymerase[J].Nat Biotechnol,2012,30(4):349-353.
[7] Zhou W Y, Tang D S , Yu-Bao Li , et al. Self-organized formation ofhexagonal Namopore arrays in anodic alumina [J]. Chinese Physics:English Version, 2001.
[8] Yuan Z,Wang C,Yi X,etal. Solid-statenanopore[J]. Nanoscale Res Lett,2018,13(1):56.
[9] Tian Li, Zhang Ying, Zhao Yunfeng. Development and application of next-generation sequencing technologies [J]. Biotechnology Bulletin, 2015,31 (11): 1-8.
[10] Zhang Xiaozhen, especially Chongge. New advances in next-generation gene sequencing technology [J]. Journal of Lanzhou University (Medical edition), 2016,42 (3): 73-80.
[11] Zhuangzi, Meng Yutong, Liu Runyang, etc. Nanopore sequencing technology and its application in pathogen diagnosis [J]. Journal of Jiangsu University (Medical edition), 2023,33 (6): 502-508
[12] Liu Kejun, Guo Shifu, Cui Le, etc. A comparative study on the application of gene sequencing technology in the clinical testing field and the regulatory status at home and abroad [J]. China Pharmaceutical Corporation, 2018,32 (11): 1520-1530
Our companys product recommendation:
1.179552-75-1 https://www.bicbiotech.com/product_detail.php?id=5500
2.197892-69-6 https://www.bicbiotech.com/product_detail.php?id=5501
3.83249-04-1 https://www.bicbiotech.com/product_detail.php?id=5502
4.96036-02-1 https://www.bicbiotech.com/product_detail.php?id=5503
5.1629249-33-7 https://www.bicbiotech.com/product_detail.php?id=5504




