This article is reprinted from the journal Clinical Laboratory by Gu Dequan, Chen Rongrong, Gao Rui, Su Jianrong
The incidence and mortality rate of infectious diseases are among the top 10 in the world, posing a serious threat to human life and health [1]. Identification of pathogens from clinical samples is key to guiding the treatment and management strategies of infectious diseases. At present, a variety of traditional diagnostic techniques are used in pathogen diagnosis, including microbial culture, detection of pathogen-specific antigen antibodies, and molecular identification of microbial nucleic acids (DNA or RNA) [2]. However, with the wide variety of pathogenic microorganisms, low flux of clinical traditional methods and time-consuming traditional culture methods, pathogen diagnosis is still difficult in about 50% of infected patients [3]. The application of high-throughput sequencing technology, also known as next-generation sequencing technology (Next Generation Sequencing, NGS) in the detection of clinical pathogens shows great potential. Among them, the main methods for identifying clinical pathogenic microorganisms based on NGS are: metagenomic sequencing (metagenomics next-generation sequencing, mNGS) and targeted sequencing (targeted NGS, tNGS). The mNGS technology is an unbiased, preset nucleic acid sequences of all potential pathogens (bacteria, viruses, fungi and parasites) from samples through the second / third generation sequencing platform. The tNGS technology is a sequencing technology based on NGS targeted enrichment (Target enrichment), which enriches tens to hundreds of known pathogenic microorganisms and their virulence and / or drug resistance genes through super-multiplex PCR amplification or hybrid capture technology, and then conducts high-throughput multipathogen parallel detection based on NGS. This paper will analyze the value and limitations, advantages and disadvantages of mNGS and tNGS technology in the detection of pathogenic microorganisms.
1、Value and limitation of mNGS technology in the detection of pathogenic microorganisms
To date, multiple studies have confirmed the promise of mNGS in clinical and public health. In 2013, Professor C.Y.Chius team successfully used mNGS technology for the first time to identify Leptospira [4] in a sample of boys with fever of unknown cause. Accurate etiology diagnosis brought accurate antimicrobial treatment, and finally the patient recovered smoothly. In 2019, Blauwkamp et al explored the detection of mNGS for sepsis patients, and identified 18.1% microorganisms in blood culture, while mNGS detected 48.6%, and maintained the sensitivity of 92.9% and 62.7% specific [5], and the consistency with blood culture reached 93.6%. The Precision Diagnosis of Acute Infectious Diseases (PDAID) study recruited 204 patients with idiopathic meningitis, encephalitis or myelitis, and found that the negative consensus rate of mNGS and conventional testing was 98%, and mNGS increased the diagnosis of the disease by 22% [6]. An etiologic study in children with unexplained fever confirmed that identification of the etiology by mNGS avoids the use of unnecessary antimicrobial [7]. To sum up, the current mNGS in respiratory tract infection, bone and joint infection, complex and atypical infection and immune suppression and other special population is widely used in [3], for unexplained fever, recurrent infection, nervous system infection, blood stream infection and eye infection etiology identification provides the solution [8,9], also can be used as a acute critical infection patients, rare pathogen infection pathogen screening technology [10].
1. Value of mNGS technology in clinical pathogenic microorganism detection: there is no need to cover a wide range of pathogenic microorganisms to facilitate the identification of new, rare and difficult to detect pathogenic agents.
(1) Traditional testing methods can only detect known pathogens, with limited detection throughput and high dependence on the experience of clinicians, which makes the diagnosis of new-onset / rare pathogen infection a challenge. The mNGS enables extensive unbiased pathogenic microbial detection by sequencing all nucleic acids in a sample. This provides the possibility of identifying clinically unknown infection and rare pathogenic bacteria. For example, in December 2019, severe pneumonia of unknown cause occurred in Wuhan, Hubei Province, China. The novel coronavirus (SARS-CoV-2) was identified as the cause of pneumonia by mNGS [11]. Patients with chlamydia parrot fever will have different degrees of pneumonia, but traditional detection of chlamydia parrot fever is time consuming and low sensitivity, so it is difficult to diagnose such pathogens. Gu et al recently reported five cases of Chlamydia psittacaris pneumonia diagnosed by mNGS method, allowing patients to quickly benefit from targeted antimicrobial therapy [12]. In addition to viruses, there are still some pathogens that are difficult to culture or need a long culture time, which are difficult to detect pathogens under traditional methods. As a culture-independent test, mNGS enables the detection of these refractory pathogens in a shorter time period. Miao et al have evaluated the clinical practice performance of mNGS in a cohort of 561 patients with acute or chronic infection, and the study confirmed a sensitivity of 50.7% and a specificity of 85.7% for mNGS. Moreover, mNGS has a better ability to detect bacteria than culture, especially for Mycobacterium tuberculosis, viruses, anaerobes and fungal [13]. A study assessing the feasibility of mNGS to detect M. tuberculosis showed that mNGS took only three days to identify 67.23% (80 / 119) of infected cases, while only 49.58% (59 / 119) of infections were detected by traditional methods, and some required more than 90 days of culture. Meanwhile, mNGS showed similar sensitivity to Xpert and conventional cultures to detect M. tuberculosis (47.92% vs 45.83%, 46.81%) [14].
(2) MNGS detection of patients treated with antibiotics, mixed infection patients have advantages: empirical use of antibiotics is still common in the treatment of patients with acute severe infection, but the problem is that it may affect the sensitivity of traditional methods (such as culture) identification of pathogens [15]. Since the mNGS technique does not rely on live bacteria but is based on microbial nucleic acids, it is relatively less affected by exposure to previous antimicrobial agents. In a study of a cohort of 561 patients with acute or chronic infections, Miao et al found that the sensitivity of mNGS in patients treated with antimicrobial therapy was significantly higher than in culture (52.7% vs 34.4%), while mNGS findings in patients with and without antimicrobial therapy were similar [13]. A study of mNGS for pathogen detection of suspected central nerve infection showed that some pathogens could still be detected by mNGS after antimicrobial treatment. The sensitivity of mNGS in patients with antimicrobial treatment was 66.67%, which was slightly lower than the sensitivity of 90% in patients without antimicrobial treatment [16]. Infections in immunocompromised people are often more complex, and the consequences of infections are more severe. Immunocompromised patients are more likely to develop mixed infection, and some studies have confirmed that mNGS has a clear advantage in the diagnosis of mixed infection. A retrospective study evaluating the detection performance of mNGS based on 55 patients with a mixed lung infection, found that mNGS had higher sensitivity than conventional testing (97.2% vs 13.9%) but lower specificity than conventional testing (63.2% vs 94.7%) [17].
2. Limitations of mNGS widely used in pathogen detection: mNGS has unique advantages for clinical microbial detection, which has been relatively popular in clinical application, but its ability to detect some demanding / difficult / low load microorganisms still needs to be improved, and the interpretation of mNGS results caused by broad microbial coverage also has great challenges.
(1) The ability of mNGS to detect some demanding / difficult / low load microorganisms also needs to be improved: mNGS can achieve extensive and unbiased detection of pathogenic microorganisms by sequencing all nucleic acids in the sample. Microbial sequence number (reads) is the key to support the correct identification of pathogenic microorganisms, sequence number refers to the number of short DNA sequences detected, a specific pathogen sequence number and the number of microorganisms in the sample, but also by the sequencing data, the size of the pathogen genome, pathogen genome specific sequence proportion, pathogen nucleic acid extraction difficulty and human host nucleic acid proportion. Since the human genome is far larger than the microbial genome and the number of host cells in samples is usually much higher than that of microbial cells, the proportion of host nucleic acid is the most important factor affecting the correct identification of pathogenic microorganisms in mNGS. The nucleic acid proportion of the host varies between different samples, ranging from 80% to 99% [8,18]. Removal of host nucleic acids is a technical solution currently being explored to improve the detection capacity of mNGS. There are currently various methods to remove / reduce host nucleic acids in samples, including physical methods such as centrifugation or filtration [19,20], chemical methods such as differential lysis of chemical reagents [21] and [22] by selective hybridization removal based on CRISPR-Cas 9 method. However, these methods will lose some of the pathogen [23] to some extent. Furthermore, the use of additional reagents during host removal may increase the risk of exogenous background contamination [2]. The complexity of the host cell proportion of different samples also increases the difficulty of dehost nucleic acids. There is no more effective and feasible method and standard for removing host nucleic acids.
In addition to the host nucleic acid proportion factor, the pathogen itself characteristics such as small genome or low load and high pathogenic microorganisms are also important factors affecting the detection performance. In a study to detect viruses based on mNGS, it was found that the supporting reads of 10 viruses detected by mNGS method were below 10, and the average genome coverage of the virus was about 2% [24]. At such low genome coverage, the number of supporting reads becomes important. However, some viruses have limited load. In a study of pediatric encephalitis astrovirus, only 1600 (0.0012%) of astrovirus reads [18] at mNGS sequencing data reached 134M. In addition, RNA viruses such as human immunodeficiency virus, Zika virus and respiratory syncytial virus also have load problems. Due to the similarity between some retroviruses and human transcription elements, the failure to distinguish the two with fewer reads leads to the failure to accurately report [25].
In addition, some pathogens with thicker cell walls, such as fungi and intracellular bacteria, may reduce the input microbial nucleic acid load due to the lower nucleic acid extraction efficiency, affecting the detection performance [18]. Based on this, there are still some controversies in the application of mNGS in fungal detection. Wang et al. analyzed 21 patients with fungal pneumonia by mNGS and detected 19 patients by mNGS, but the study found that mNGS did not show a dominant [26] in the detection of cryptococcus. Another study by Fang et al. also suggested that mNGS has its advantages and disadvantages in fungal detection. The traditional detection methods had good detection performance of C. albicans and Candida tropicalis, while mNGS had better detection performance of Aspergillus [27]. In another study of mNGS to detect patients with meningitis, although mNGS and traditional tests showed high consistency, there were missed and missed patients. M. bovis, M. tuberculosis, and Cryptococcus neoformans were detected in the mNGS of the missed patients, but the abundance did not reach the predetermined reporting threshold [28]. In order not to detect some pathogenic microorganisms, such as Mycobacterium tuberculosis and Legionella, these microorganisms generally set a low threshold [24,25]. However, at such a low threshold, it can also make clinical judgment and decision-making difficult to [10].
(2) The wide spectrum of microbial coverage leads to mNGS results interpretation is a huge challenge: mNGS can theoretically detect all currently known pathogens with about 8,000 or even currently unknown genome sequences. Therefore, in the mNGS test results, microbial contamination in the environment, reagents, containers, as well as a large number of non-pathogenic microorganisms, and pathogenic microorganisms such as colonization in the human body are mixed together [29]. This requires not only a database containing all pathogenic microorganisms but also a relevant database for mNGS analysis for filtering [30]. Considering the pathogenicity and pathogenic conditions of different microorganisms, mNGS needs to set a personalized quote threshold of [3]. For example, the consensus of highly pathogenic but low pathogenic pathogens such as Mycobacterium tuberculosis, Yersinia pestis and epidemic hemorrhagic fever virus suggests that one sequence should be considered positive even on the premise of excluding contamination [13,31-33]. Even so, the number of microorganisms reported by mNGS is 5-20, which makes the clinical interpretation of mNGS extremely challenging, requires a lot of clinical, pathogenic microbial diagnosis and professional experience [3], need to consider the overall clinical situation [5], which usually requires a professional multidisciplinary team to complete [34].
2、Advantages and disadvantages of tNGS technology in the detection of pathogenic microorganisms
The tNGS technology is a sequencing technology based on NGS targeted enrichment (Target enrichment). In the detection of pathogens, dozens to hundreds of known pathogenic microorganisms (especially low concentrations) and their virulence and / or drug resistance genes were enriched in the samples to be tested by ultra-multiplex PCR amplification or hybridization capture technology, and then high-throughput parallel detection based on NGS. Targeted enrichment is a key link in tNGS technology. Through the [35] enrichment of the target genomic regions (regions of interest, ROI), the ROI regions have sufficient sequencing depth and coverage, and then successfully identify the target pathogens. Relative to mNGS technology, tNGS can enrich microbial nucleic acids by tens or even tens of thousands of times. At present, the targeted enrichment strategies of tNGS technology at home and abroad are mainly divided into two types: tNGS based on super-multiplex PCR amplification and tNGS [35] based on hybridization capture. TNGS technology has been widely popular in the field of disease detection, especially in the field of tumor gene detection, but its application in pathogenic microorganism detection is still in its infancy, and related research literature is few. This part mainly expounds tNGS targeted enrichment strategy, technical advantages, technical difficulties and other aspects.
1. TNGS targeted enrichment strategy: Among the two main targeted enrichment methods of tNGS, hybridization capture targeted sequencing mainly uses probe hybridization to enrich target fragments, which is suitable for the comprehensive detection of target regions of the genome. At present, the application is mainly liquid phase hybridization capture sequencing, that is, based on the principle of base complementary pairing, design and synthetic nucleic acid probes, DNA libraries for hybridization enrichment of target regions based on liquid phase environment, and sequencing. For hybridization capture sequencing, it involves many technical development difficulties, such as probe sequence design, selection of target pathogen, and construction of refined database. In probe design, we need to evaluate the sequence characteristics of the coverage position, and if many of the probes fall in the repetitive sequence region, or the high-copy sequence region, the probe will bind more non-target regions. Design more specific probes can effectively reduce the binding of non-specific sequences and improve capture specificity. In addition, the coverage of the pathogen genome probe, whether the pathogen genome section can be evenly identified; whether the probe can accurately identify / distinguish closely related species, how the coverage of the specific sequence marker; how different pathogen subtypes (strains) should be considered comprehensively.
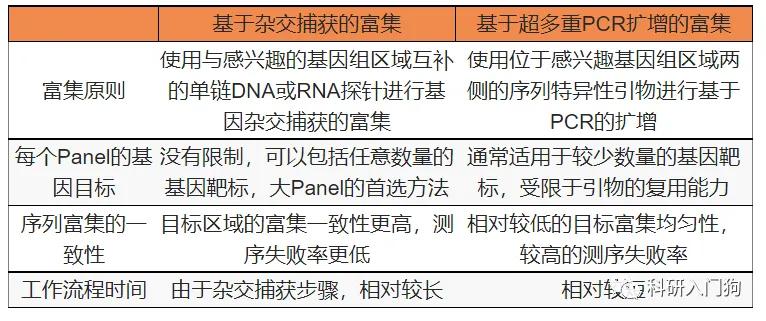
Super-multiplex PCR targeted sequencing is the technology of designing multiplex PCR primers for amplification and enrichment and sequencing for the target region of interest. Usually applicable to detecting tens to thousands of sites, or regions under tens of kb. Its core is the primer design, through PCR extension enrichment target segment, and then library construction, suitable for the target area is relatively small, for low copy number of template DNA, can produce enough number for sequencing amplicons, this method can obviously improve the efficiency, save time, reduce economic cost, the difficulty is that multiple interference between primers and non-specific amplification. Studies have also shown that the hybridization capture-based tNGS technology has better uniformity than the amplicon-based methods, and may have more objective true content of pathogens in reaction samples [26], but the technology development is more difficult and more complicated detection process than the super multiplex PCR amplification method. The main differences between the two methods are shown in Table 1.
Table 1. Main differential comparison of enrichment methods for hybridization capture targeted sequencing and super-multiplex PCR targeted sequencing
2.Technical advantages of tNGS: Several studies compared mNGS with tNGS, which also confirmed the feasibility of clinical application of tNGS. A recent comparative study of mNGS and tNGS in BALF samples from patients with respiratory tract infections showed that tNGS showed similar microbial detection limits to mNGS at only one-third of mNGS (limit-of-detection, LoD). The study covered the tNGS detection of 280 pathogenic microorganisms, and its performance was comparable to that of mNGS. The accuracy of tNGS was 65.6%, while mNGS was 67.1% [36] in the same cohort. Similarly, in another report, tNGS covering more than 150 microorganisms showed mNGS a similar detection rate (82.17% vs 86.51%), and the detection rate were also consistent in bacteria, fungi and viruses [37].
(1) Enrichment increases the coverage of target microorganism detection to improve the detection credibility: tNGS technology can specifically target the enrichment of target pathogen nucleic acid sequences, specifically increase the number and proportion of target pathogen sequences in the sequencing data, and improve the number of reads and coverage of target microorganism detection by [2]. Targeted enrichment of tNGS, even under conditions without host removal, can specifically increase the proportion of microbial nucleic acids. One study compared the improvement of mNGS virus detection ability by host nucleic acid removal and virus targeted capture enhancement, and found that the load of viral nucleic acid could be increased by 90%, but the effect of host nucleic acid removal method fluctuated greatly [38]. In a study by Zhao et al., who compared tNGS with mNGS based on ribosomal RNA enrichment, they found that the proportion of microbial nucleic acids in the sample increased by 7-fold, whether viral or fungal or drug resistance genes, and the detection results of tNGS and clinical consistency were high [25]. After enrichment, tNGS can improve the load and coverage of caustic and low-load pathogens, and improve the credibility of detection. Comparing differences in detecting viruses based on mNGS and against virus capture, Todd et al. The study found that in the samples of children with unexplained fever, the number of viral detected virus increased 674 times, and the coverage of the viral genome increased from 2.1% to 83.2%. Thanks to the improved coverage, the captured tNGS can be used to analyze the [24] of cycloviruses with genome variation of up to 58%. In another study comparing the test results of tNGS and mNGS in patients with pulmonary infection, more mixed infections of B. tuberculosis were detected in tNGS, double [37] more than in mNGS. In conclusion, the targeted enrichment of tNGS can increase the number of corresponding sequences corresponding to microorganisms and contribute to the credible detection of microorganisms.
(2) TNGS can stably carry out the detection of drug resistance or virulence genes: the development of sequencing technology makes people have a deep understanding of the genotype, virulence, drug resistance genes and systemic genetic background of microbial strains. Analysis of the drug resistance gene data of 1379 S. aureus isolates by whole genome sequencing, Mason et al. And found that the correlation between resistance genotype and phenotype could be as high as 98.3% [39]. In the detection of drug resistance genes, for the representative drug resistance of ecA, KPC and other genes, it is usually necessary to identify whether there is drug resistance type in the infected samples, but the difficulty is how to combine the drug resistance genes with the pathogen, it is necessary to interpret drug resistance and virulence genes; on the other hand, the representative drug resistance of tuberculosis needs to be accurate to the mutation site [40]. The metagenomic sequencing pathogenic microorganisms detection bioinformatics analysis standardization management expert consensus, points out that mNGS because of the microbial sequencing sequence of 5% of the total sequence, specific sequence coverage less than 1% of the microbial genome, in this case drug resistance genes, virulence island gene, transcriptome and metabolic pathway analysis is almost impossible [32]. Therefore, the mNGS technology for drug resistance and virulence gene detection has limited [2].
The tNGS technology can target region enrichment of resistance / virulence genes, ensuring accurate and stable detection. In an exploratory study based on 201 tNGS from respiratory samples for the analysis of pathogenic microorganisms and drug resistance gene testing, 53.8% of drug resistance-related genes reported by tNGS were consistent with clinical susceptibility testing [36]. However, it is necessary to consider that the drug resistance effect of the same drug resistance gene varies in different microorganisms. In the above studies, multiple P. aeruginosa strains carrying carbapenemase-related OXA resistance genes were found, but the resistance phenotype [36] of these microorganisms was not found in clinical drug susceptibility analysis. Therefore, when targeting target resistance / virulence genes, different mutation types of the same gene should be considered to produce different resistance effects, and achieve full coverage of resistance / virulence genes.
(3) Due to the clear setting range of the target pathogen, tNGS detection can significantly simplify the interpretation process: In the comparative study of mNGS and tNGS, it was found that the number of pathogens reported in a single time based on mNGS detection sample was 4-24, while tNGS detection was concentrated in 1-2 [36]. In terms of the construction of pathogenic microorganism database, compared with the "large and complete" of tens of thousands of microorganisms in mNGS database, tNGS database can focus more on dozens to hundreds of target pathogens, and more deeply and meticulously distinguish between different subspecies and subtypes. The study showed that in the automatic birth letter analysis phase, 96.8% of the tests in mNGS were interpreted as background microorganisms, compared with 75.2% [36] in tNGS. Even so, the fine design of the tNGS database is complex and necessary. The fine design of the database can ensure the accurate screening of pathogenic microorganisms within the targeted detection range, especially for the coverage of rare and demanding microorganisms, and for the fine identification of common pathogenic bacteria to the species level. In the comparison of tNGS and mNGS studies, both methods detected Pneumostiia carinii (P. carinii), but considering that it was not a human pathogenic organism. However, C. neoformans detected in tNGS was misidentified as Cryptococcus [36]. This suggests that the refined, specialized design of the tNGS birth database is crucial in pathogen identification.
3. Lack of clinical application of tNGS: The detection panel of tNGS is designed according to known pathogens, so tNGS is not suitable for the detection of new pathogenesis. Although panel can theoretically include dozens or even hundreds of pathogens, panel may still need to be designed in combination with clinical application scenarios. If the infectious pathogen itself is not within the design range of Panel, it cannot be detected. In the recent comparative study of tNGS and mNGS, a large number of human herpesviruses were detected in mNGS, but the pathogen detection range of tNGS did not include the virus in tNGS; Besides, mNGS also detected 8 bacteria, 2 parasites, 5 viruses and 5 fungi [37] outside the detection range of tNGS.
3、Outlook for the clinical application of mNGS and tNGS
Through metagenomic sequencing of clinical specimens, mNGS technology can detect a variety of pathogenic microorganisms (including viruses, bacteria, fungi and parasites) in the specimens without bias. More and more clinical studies and case reports of special pathogen infection affirm the value of mNGS in the auxiliary diagnosis of infectious diseases. At present, mNGS is mainly used in patients with acute and severe diseases suspected of infection, negative routine tests but suspected difficult and severe infections. Given the price and interpretation difficulties, mNGS is not suitable for initial screening of routine infections. The tNGS designs panel for primary clinical screening needs to cover common pathogens, and improves the confidence of pathogen detection through targeted amplification. This technology is expected to fill the demand gap between traditional molecular detection and mNGS. Therefore, tNGS is receiving increasing attention in the field of clinical examination.
Of course, with the application of NGS technology in the precise diagnosis and treatment of infectious diseases, it is necessary to promote the technical optimization of mNGS and tNGS detection and the application standardization. At the technical level, automated process and accurate and more rapid sequencer may be the two major strengths of mNGS and tNGS technology transformation; at the application level, both mNGS and tNGS need to further clarify the clinical use scenarios, and identify the sample type, the target pathogen range, the positive judgment value, and the product matching database according to the clinical application scenario. In the future, the pathogen detection methods based on small and medium coverage of traditional detection, tNGS and mNGS will be more effective and accurate for accurate clinical diagnosis and treatment.
reference documentation
Lozano, R, et al. Global and regional mortality from 235 causes of death for 20 age groups in 1990 and 2010: a systematic analysis for the Global Burden of Disease Study 2010. Lancet, 2012. 380(9859): p. 2095-128.
Chiu CY,Miller SA. Clinical metagenomics. Nat Rev Genet, 2019. 20(6): p. 341-355.
Li N, et al. High-throughput metagenomics for Identification of pathogens in the clinical settings. Small Methods, 2021. 5(1): p. 2000792.
The companys product recommendation:
1.161040-25-1 https://www.bicbiotech.com/product_detail.php?id=5505
2.2195389-90-1 https://www.bicbiotech.com/product_detail.php?id=5506
3.2241432-85-7 https://www.bicbiotech.com/product_detail.php?id=5507
4.2361327-06-0 https://www.bicbiotech.com/product_detail.php?id=5508
5.104700-86-9 https://www.bicbiotech.com/product_detail.php?id=5509




