Targeted protein degradation (TPD) has emerged as one of the most revolutionary technological directions in recent drug development. Degradation technologies represented by PROTACs have overcome the limitations of traditional small-molecule inhibitors, achieving the therapeutic goal of "clearing pathogenic proteins." However, conventional PROTACs are highly dependent on E3 ubiquitin ligases, presenting bottlenecks such as ligand scarcity, limited applicable targets, and poor drugability.
Recently, a Spanish research team published a groundbreaking study titled "Expanding the targeted protein degradation approach with small molecule chimeras directed to the 26S proteasome" in Nature Communications, successfully developing 26S-targeted ubiquitin-independent degraders (26S-UIDs). These agents bypass E3 ligases and directly deliver target proteins to the 26S proteasome for degradation, opening a new frontier in the field of targeted protein degradation.

Core limitations of traditional PROTACs
Classic PROTAC molecules function as "molecular bridges," with one end binding to the pathogenic target protein and the other recruiting E3 ubiquitin ligases to ubiquitinate the target protein, which is ultimately recognized and degraded by the 26S proteasome. Although PROTACs have demonstrated clinical potential, their inherent limitations remain difficult to overcome:
- E3 ligands are critically scarce: The limited availability of E3 ubiquitin ligands for PROTAC construction significantly restricts molecular design possibilities.
- Dependent on the ubiquitination process: requires the target protein to possess modifiable lysine sites, as a large number of proteins lacking suitable sites cannot be degraded.
- Limited tissue specificity: The expression levels and activity of E3 ligase vary significantly across different tissues, resulting in suboptimal degradation efficiency for certain targets.
- Challenges in drug development: PROTACs exhibit large molecular weights and poor solubility, making oral absorption and in vivo metabolism difficult to optimize.
These challenges have prompted the academic community to explore novel degradation strategies that are independent of E3 ubiquitination and ubiquitination.
26S-UIDs: A Disruptive Novel Degradation Mechanism
The research team broke away from conventional approaches by selecting USP14, a 26S proteasome regulatory protein, as the recruitment target and constructed a novel small-molecule degradation system termed 26S-UIDs, whose core mechanism is entirely distinct from PROTAC:
Bispecific binding: The small molecule chimeric molecule binds to the target protein at one end and specifically binds to USP14 at the other end.
Direct targeting: Precisely transporting the target protein to the ATP enzyme motor of the 26S proteasome.
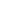
Non-ubiquitin-dependent: Does not require E3 ligase involvement or ubiquitination modification, directly triggering proteasomal unfolding and degradation of target proteins. USP14 is a 26S proteasome regulatory factor located at the base of the proteasome regulatory particle, adjacent to the ATPase channel, serving as an ideal anchoring site. Based on optimization of the USP14 small molecule ligand IU1 family, the research team screened out ligand 317 with high affinity and low inhibitory activity, significantly enhancing degradation efficiency.
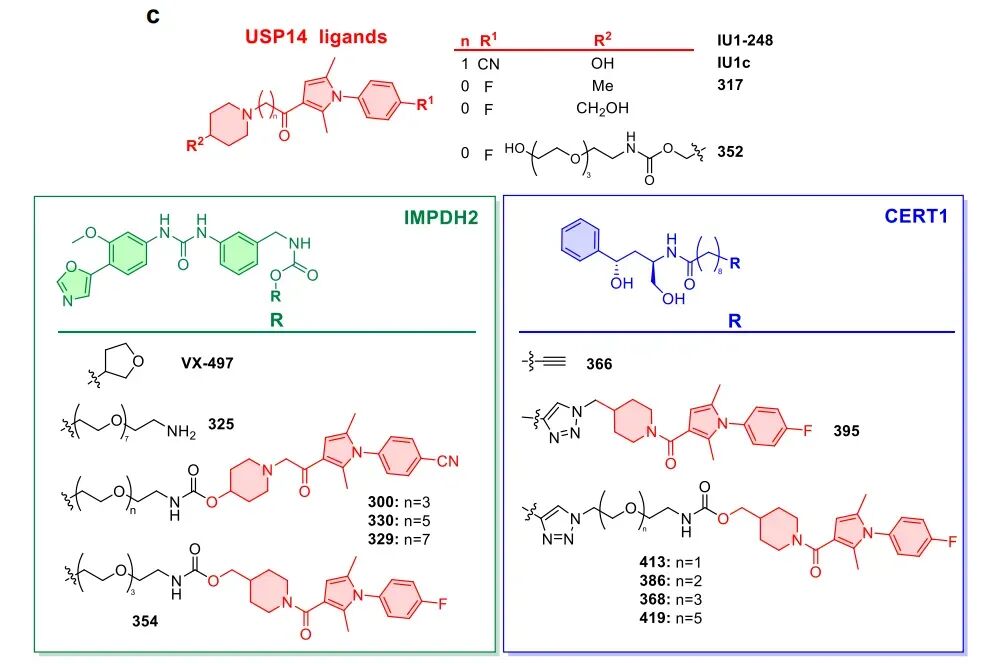
Validation of two major anticancer targets: high efficacy, specificity, and well-established mechanisms
The study selected IMPDH2 (a key target for leukemia and lymphoma) and CERT1 (a regulatory target for breast cancer drug resistance) as models to conduct systematic functional validation, with results demonstrating strong persuasiveness.
IMPDH2 is the rate-limiting enzyme for de novo guanosine nucleotide synthesis, which is highly expressed in leukemia, lymphoma, and various solid tumors, and is closely associated with tumor progression and chemotherapy resistance. Currently, only a few degradation agents have been reported. Researchers targeted the IMPDH2 inhibitor VX-497, using USP14 ligand IU1-248 and its derivatives as recruitment ends, and constructed a series of chimeras through PEG linkers of varying lengths. Cell experiments demonstrated that the optimized molecule 354 achieved complete IMPDH2 clearance at concentrations of 500–1000 nM, with significantly superior degradation efficacy compared to earlier molecules. Mechanistic studies confirmed that IMPDH2 degradation could be reversed by proteasome inhibitors but was unaffected by ubiquitin activator inhibitor MLN7243; in USP14 knockout cells, the degradation effect was completely abolished. These results directly demonstrate that 26S-UIDs achieve target protein clearance through USP14-dependent, ubiquitin-independent, and proteasome-dependent mechanisms.
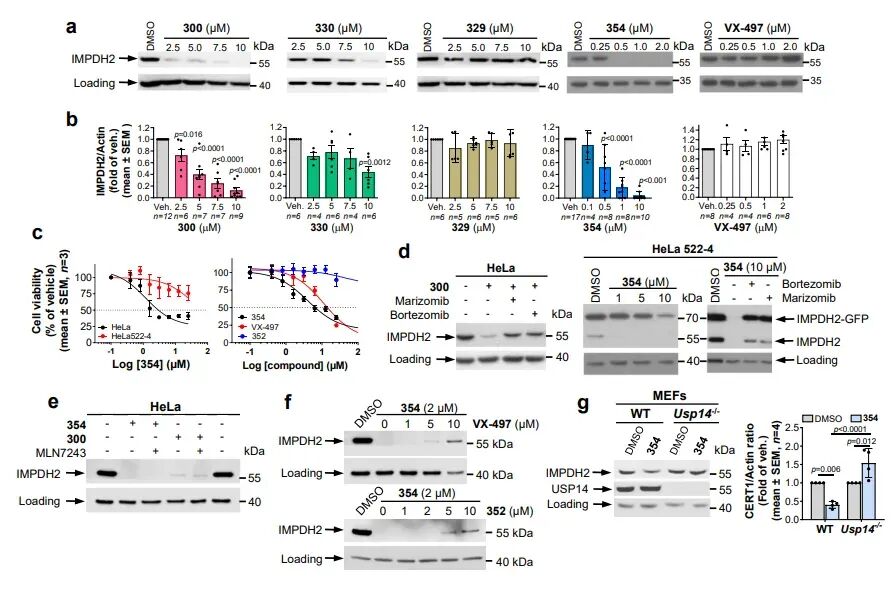
CERT1 is a ceramide transporter responsible for transporting ceramides from the endoplasmic reticulum to the Golgi apparatus for sphingomyelin synthesis. Its high expression reduces intracellular pro-apoptotic ceramide levels, thereby mediating chemotherapy resistance in tumors such as breast cancer. Prior to this study, no degradation agents targeting CERT1 had been reported. This research employed an HPA family antagonist of CERT1 as the target compound and constructed five 26S-UIDs using USP14 ligand 317. Experimental results demonstrated that the optimal molecule 368 could significantly degrade CERT1 within 16–24 hours, with a degradation mechanism identical to that of IMPDH2: dependent on 26S proteasome, dependent on USP14, and independent of ubiquitination. Lipidomics analysis confirmed that CERT1 degradation leads to significant accumulation of pro-apoptotic ceramides with C14–C20 chain lengths in cells and enhances the sensitivity of HER2-positive breast cancer cells to lapatinib, suggesting this strategy holds significant potential for overcoming tumor resistance.
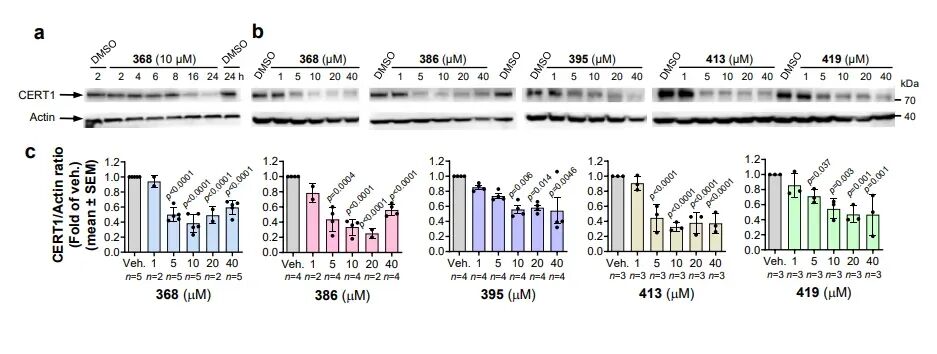
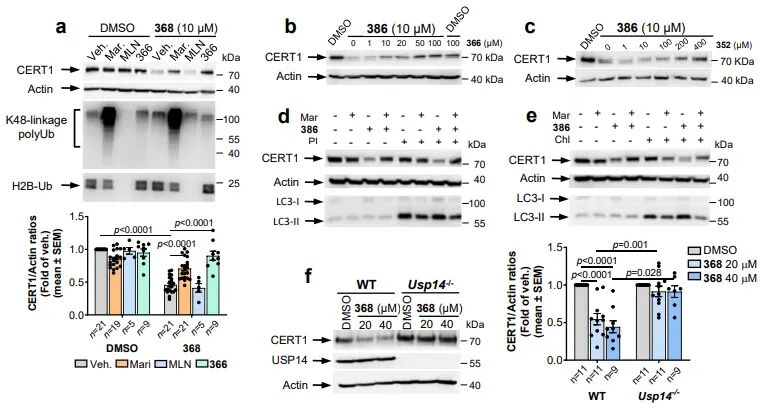
Triad Complex Validation
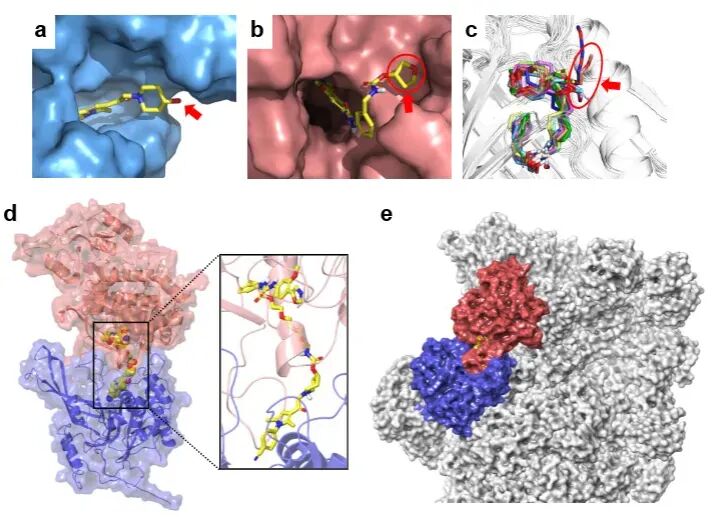
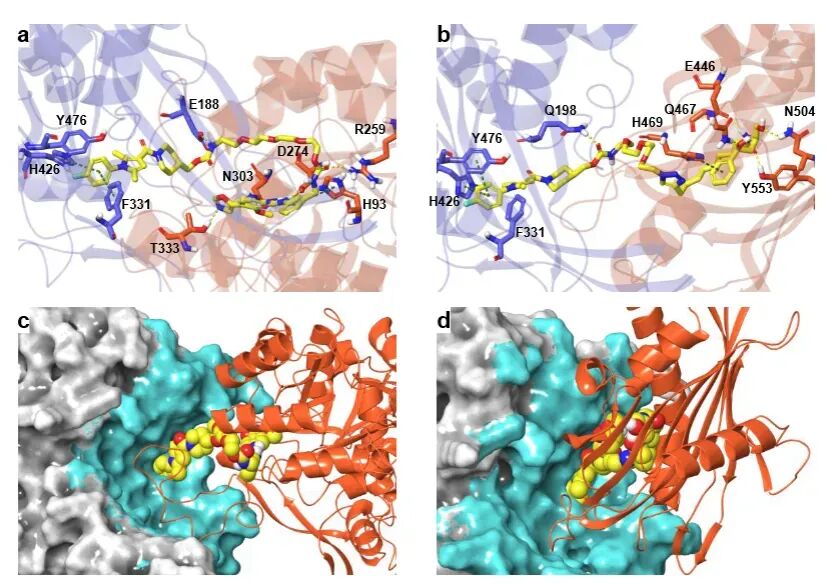
To further elucidate the molecular mechanisms, the research team confirmed through surface plasmon resonance (SPR), pull-down experiments, and computer simulations that 26S-UIDs can mediate the formation of the USP14-chimeras-target protein ternary complex. The spatial conformation of this complex is fully compatible with the structure of the 26S proteasome, without causing steric hindrance.
Unlike traditional PROTACs, the degradation efficiency of 26S-UIDs is not entirely dependent on linker length and interfacial energy, demonstrating a more flexible molecular design space.
Core advantages of 26S-UIDs technology
Overall, this study successfully established a novel targeted protein degradation platform that is independent of E3 ubiquitin ligases and substrate ubiquitination. 26S-UIDs directly recruit the 26S proteasome in small molecule form, overcoming the core limitations of traditional PROTACs and offering advantages such as superior drugability, broader target applicability, and novel mechanisms of action. The successful validation of IMPDH2 and CERT1 not only provides new candidate molecules for tumor therapy but also demonstrates the high versatility of this strategy, which can be extended to the degradation of more pathogenic proteins in the future.
Disclaimer: This article is intended solely for knowledge exchange, sharing, and popular science purposes, and does not involve commercial promotion. It should not be construed as medical guidance or medication recommendations. For copyright infringement inquiries, please contact us for removal.
Product recommendations for our company:
1.389628-28-8 https://www.bicbiotech.com/product_detail.php?id=6527
2.1071608-51-9 https://www.bicbiotech.com/product_detail.php?id=6528
3.193354-13-1 https://www.bicbiotech.com/product_detail.php?id=6529
4.102331-54-4 https://www.bicbiotech.com/product_detail.php?id=6530
5.175281-76-2 https://www.bicbiotech.com/product_detail.php?id=6531




